Sample name: K562_PRO-seq_100
Looper stats summary
| sample_name |
K562_PRO-seq_100 |
| sample_desc |
Unsampled K562 PRO-seq |
| treatment |
none |
| protocol |
PRO |
| organism |
human |
| read_type |
SINGLE |
| umi_len |
0 |
| read1 |
['/project/shefflab/data//sra_fastq/SRR1554311.fastq.gz', '/project/shefflab/data//sra_fastq/SRR1554312.fastq.gz'] |
| srr |
SRR155431[1-2] |
| Raw_reads |
496581677 |
| Fastq_reads |
496581677 |
| Reads_with_adapter |
423059183.0 |
| Uninformative_adapter_reads |
12437983.0 |
| Pct_uninformative_adapter_reads |
2.5047 |
| Trimmed_reads |
484143694 |
| Trim_loss_rate |
2.5 |
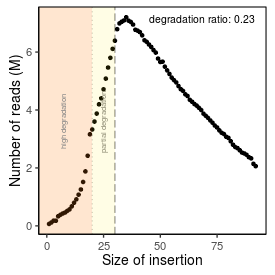
| Peak_adapter_insertion_size |
34 |
| Degradation_ratio |
0.2312 |
| Aligned_reads_human_rDNA |
44680364.0 |
| Alignment_rate_human_rDNA |
9.23 |
| Mapped_reads |
434069790 |
| QC_filtered_reads |
46741044 |
| Aligned_reads |
387328746 |
| Alignment_rate |
80.0 |
| Total_efficiency |
78.0 |
| Read_depth |
29.61 |
| Mitochondrial_reads |
9149071 |
| Maximum_read_length |
100 |
| Genome_size |
3099922541 |
| NRF |
0.28 |
| PBC1 |
0.54 |
| PBC2 |
3.89 |
| Unmapped_reads |
5393540 |
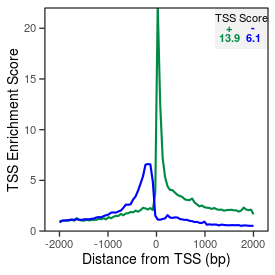
| TSS_coding_score |
13.9 |
| TSS_non-coding_score |
5.8 |
| File_mb |
38578.5 |
| Read_type |
SINGLE |
| Genome |
hg38 |
| Pause_index |
10.96 |
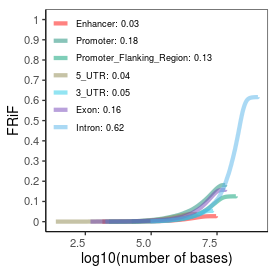
| Plus_FRiP |
0.36 |
| Minus_FRiP |
0.34 |
| mRNA_contamination |
1.37 |
| Time |
2:23:44 |
| Success |
06-14-23:34:32 |