Sample name: K562_RNA-seq_50
Looper stats summary
| sample_name |
K562_RNA-seq_50 |
| sample_desc |
50% K562 PRO-seq + 50% K562 RNA-seq |
| treatment |
70M total reads |
| protocol |
PRO |
| organism |
human |
| read_type |
SINGLE |
| umi_len |
0 |
| read1 |
/project/shefflab/data//guertin/fastq/K562_50pctRNA.fastq.gz |
| srr |
K562_50pctRNA |
| Raw_reads |
70000000 |
| Fastq_reads |
70000000 |
| Reads_with_adapter |
42246395.0 |
| Uninformative_adapter_reads |
875215.0 |
| Pct_uninformative_adapter_reads |
1.2503 |
| Trimmed_reads |
69124785 |
| Trim_loss_rate |
1.25 |
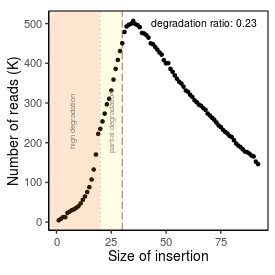
| Peak_adapter_insertion_size |
34 |
| Degradation_ratio |
0.231 |
| Aligned_reads_human_rDNA |
3959053.0 |
| Alignment_rate_human_rDNA |
5.73 |
| Mapped_reads |
59023095 |
| QC_filtered_reads |
9634021 |
| Aligned_reads |
49389074 |
| Alignment_rate |
71.45 |
| Total_efficiency |
70.56 |
| Read_depth |
6.57 |
| Mitochondrial_reads |
2741453 |
| Maximum_read_length |
100 |
| Genome_size |
3099922541 |
| NRF |
0.73 |
| PBC1 |
0.89 |
| PBC2 |
14.29 |
| Unmapped_reads |
6142637 |
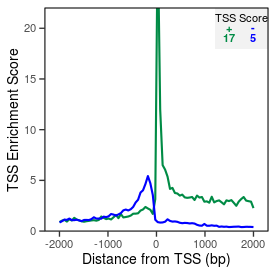
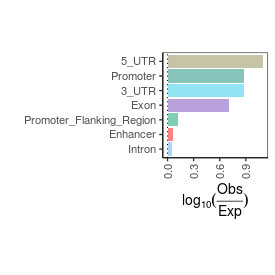
| TSS_coding_score |
17.0 |
| TSS_non-coding_score |
4.6 |
| File_mb |
5536.54 |
| Read_type |
SINGLE |
| Genome |
hg38 |
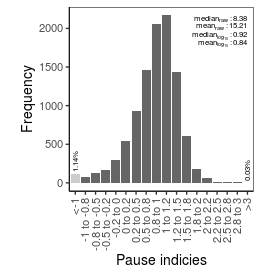
| Pause_index |
8.38 |
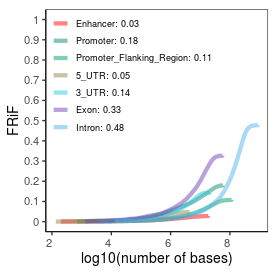
| Plus_FRiP |
0.38 |
| Minus_FRiP |
0.37 |
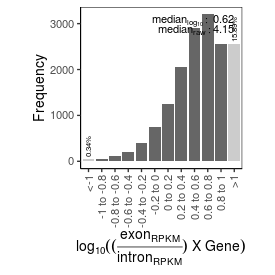
| mRNA_contamination |
4.15 |
| Time |
0:35:54 |
| Success |
06-14-21:47:41 |