Sample name: H9_PRO-seq_10
Looper stats summary
| sample_name |
H9_PRO-seq_10 |
| sample_desc |
10% subset H9 PRO-seq 2 |
| treatment |
DMSO |
| protocol |
PRO |
| organism |
human |
| read_type |
PAIRED |
| umi_len |
8 |
| read1 |
/project/shefflab/data//guertin/fastq/H9_PRO-seq_10pct_PE... |
| read2 |
/project/shefflab/data//guertin/fastq/H9_PRO-seq_10pct_PE... |
| srr |
H9_PRO-seq_10pct |
| Raw_reads |
11560826 |
| Fastq_reads |
11560826 |
| Reads_with_adapter |
4579820.0 |
| Uninformative_adapter_reads |
2857948.0 |
| Duplicate_reads |
81668.0 |
| Pct_uninformative_adapter_reads |
49.4419 |
| Trimmed_reads |
5661748 |
| Trim_loss_rate |
51.03 |
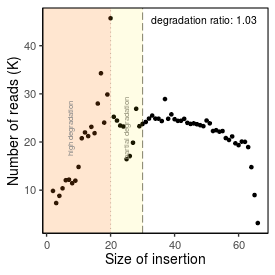
| Peak_adapter_insertion_size |
20 |
| Degradation_ratio |
1.033 |
| Aligned_reads_human_rDNA |
583234.0 |
| Alignment_rate_human_rDNA |
10.3 |
| Mapped_reads |
4366101 |
| QC_filtered_reads |
2551719 |
| Aligned_reads |
1814382.0 |
| Alignment_rate |
32.05 |
| Total_efficiency |
15.69 |
| Read_depth |
1.54 |
| Mitochondrial_reads |
83974 |
| Maximum_read_length |
38 |
| Genome_size |
3099922541 |
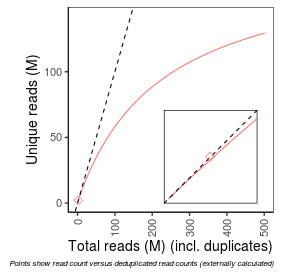
| Frac_exp_unique_at_10M |
0.8545 |
| NRF |
1.0 |
| PBC1 |
972859.0 |
| PBC2 |
972859.0 |
| Unmapped_reads |
667337 |
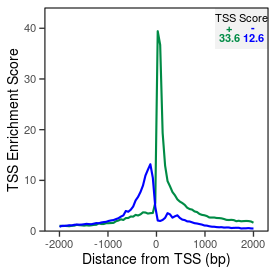
| TSS_coding_score |
33.6 |
| TSS_non-coding_score |
12.2 |
| File_mb |
249.31 |
| Read_type |
PAIRED |
| Genome |
hg38 |
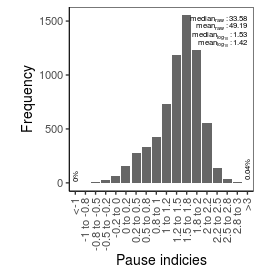
| Pause_index |
33.58 |
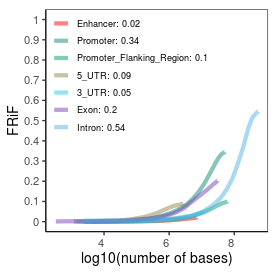
| Plus_FRiP |
0.39 |
| Minus_FRiP |
0.36 |
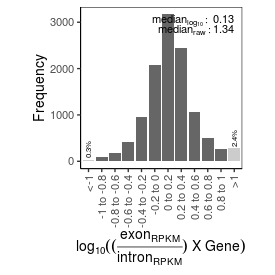
| mRNA_contamination |
1.34 |
| Time |
0:36:56 |
| Success |
06-14-21:55:30 |