Sample name: K562_RNA-seq_10
Looper stats summary
| sample_name |
K562_RNA-seq_10 |
| sample_desc |
90% K562 PRO-seq + 10% K562 RNA-seq |
| treatment |
70M total reads |
| protocol |
PRO |
| organism |
human |
| read_type |
SINGLE |
| umi_len |
0 |
| read1 |
/project/shefflab/data//guertin/fastq/K562_10pctRNA.fastq.gz |
| srr |
K562_10pctRNA |
| Raw_reads |
70000000 |
| Fastq_reads |
70000000 |
| Reads_with_adapter |
56155013.0 |
| Uninformative_adapter_reads |
1576345.0 |
| Pct_uninformative_adapter_reads |
2.2519 |
| Trimmed_reads |
68423655 |
| Trim_loss_rate |
2.25 |
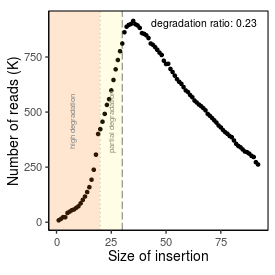
| Peak_adapter_insertion_size |
34 |
| Degradation_ratio |
0.231 |
| Aligned_reads_human_rDNA |
5831595.0 |
| Alignment_rate_human_rDNA |
8.52 |
| Mapped_reads |
60754568 |
| QC_filtered_reads |
7195486 |
| Aligned_reads |
53559082 |
| Alignment_rate |
78.28 |
| Total_efficiency |
76.51 |
| Read_depth |
5.9 |
| Mitochondrial_reads |
1579435 |
| Maximum_read_length |
100 |
| Genome_size |
3099922541 |
| NRF |
0.77 |
| PBC1 |
0.89 |
| PBC2 |
11.77 |
| Unmapped_reads |
1837492 |
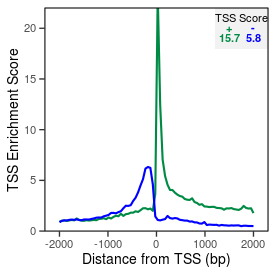
| TSS_coding_score |
15.7 |
| TSS_non-coding_score |
5.5 |
| File_mb |
5660.06 |
| Read_type |
SINGLE |
| Genome |
hg38 |
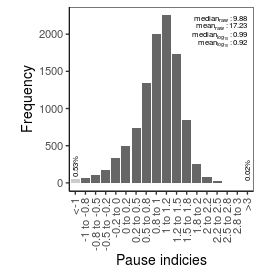
| Pause_index |
9.88 |
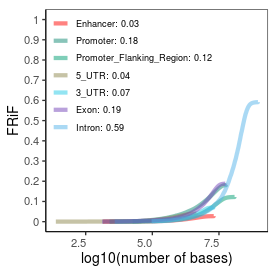
| Plus_FRiP |
0.36 |
| Minus_FRiP |
0.34 |
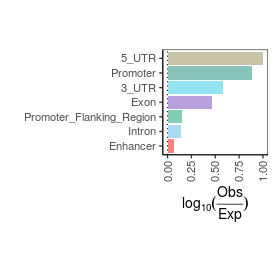
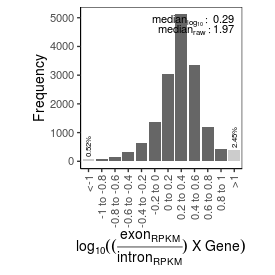
| mRNA_contamination |
1.97 |
| Time |
0:37:46 |
| Success |
06-14-21:49:02 |